Biography
I am a computational biologist focusing on biomolecules and their interaction and regulation, which form the molecular foundation of biomedical science and future in silico (virtual) biology. I am currently a postdoc with Yufeng Shen at Columbia University, where I investigate protein fitness landscapes from both evolutionary and biophysical perspectives. I developed SeqDance and ESMDance, two protein language models (pLMs) trained on protein dynamics data. I explained the scaling behavior of pLMs for fitness prediction (why larger models do not always perform better). I also developed MotifAE for unsupervised discovery of functional motifs from pLM. I received my Ph.D. in Biomedical Informatics from Peking University with Tingting Li. During my doctoral work, I studied biomolecular interactions involved in degradation regulation and protein localization mediated by phase separation.
I was born in Zhucheng, Shandong, China (Chinese name: 侯超). Outside of research, I enjoy basketball, photography, travel, and food.
Updated: April 2026
- Biological sequence → structure ensemble → function and fitness → evolution
- Representation and generative deep learning of biomolecules
- Predicting the effects of genetic variants for precision medicine
- Biomolecular interaction network and subcellular localization
Postdoc, 2023.09-
Columbia University
PhD of Biomedical Informatics, 2020.09-2023.07
Peking University
Bachelor of Medicine and Economics, 2015.09-2020.07
Peking University
Projects
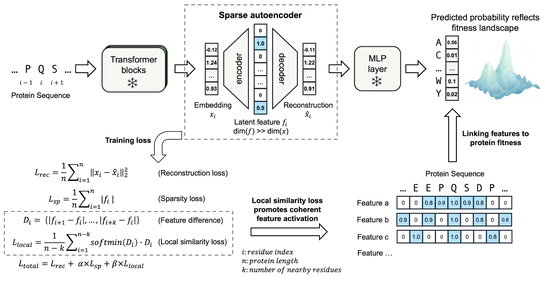
pLM interpretation
Unsupervised discovery of functional motifs from pLM
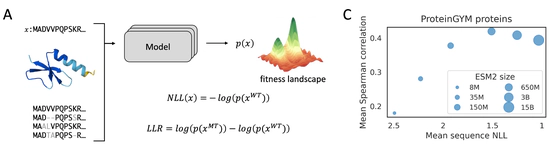
pLM fitness prediction
why larger model do NOT always perform better?
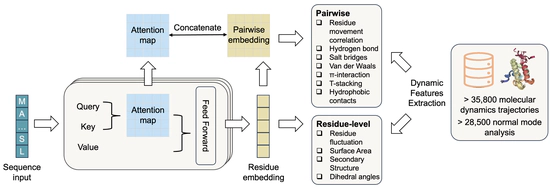
Learn protein dynamics with pLMs
SeqDance/ESMDance trained on Protein Dynamic Properties

Bioinformatics tools for phase seperation
PhaSepDB, PhaSePred and MloDisDB
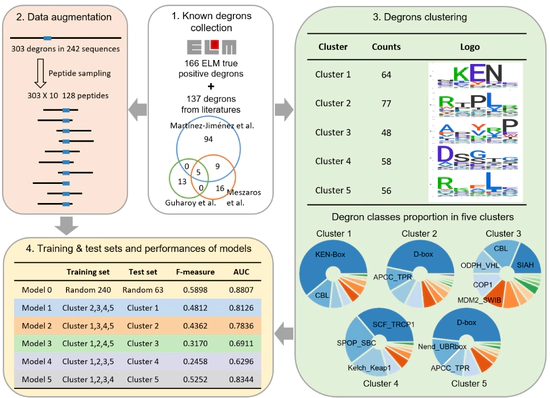
Predicting E3 ligase binding site
Degpred predicts degron and binding E3 via deep learning
Main Publications
Contact
- ch3849@cumc.columbia.edu
- 1130 St Nicholas Ave, New York, NY 10032
