MotifAE Reveals Functional Sequence Patterns from Protein Language Model: Unsupervised Discovery and Interpretability Analysis
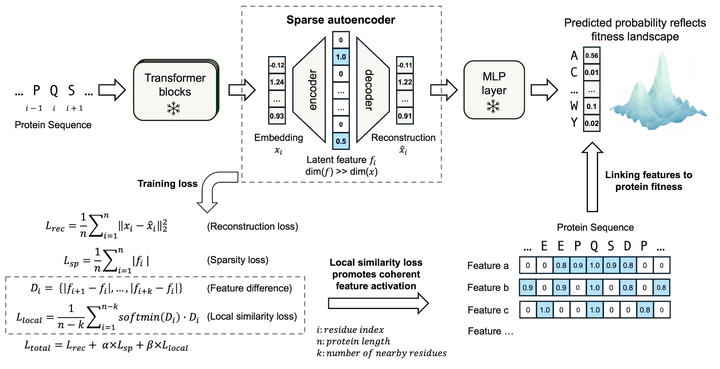 Image credit: Chao Hou
Image credit: Chao HouAbstract
Protein language models (pLMs) learn sequence patterns at evolutionary scale, but these patterns remain inaccessible within these “black box” models. To discover them, we developed MotifAE, an unsupervised framework based on the sparse autoencoder (SAE) architecture that projects pLM embeddings into an interpretable, sparse latent space. MotifAE introduces an additional smoothness loss to encourage coherent feature activation, which markedly improves the identification of known functional motifs compared to the standard SAE. The sequence patterns captured by MotifAE exhibit rich diversity, align with known functional motifs, and are reflected in the model’s weight space. Beyond short motifs, MotifAE also captures structural domains, with latent feature activation scores correlating with residue importance for different domain functions. By aligning MotifAE features with experimental data, we further identified features associated with domain folding stability. These features enable the prediction of a stability-specific fitness landscape that improves stability prediction and supports the engineering of domains with enhanced stability. Overall, MotifAE provides a general framework for systematic sequence pattern discovery and interpretation, with the potential to advance protein function analysis, mutation effect interpretation, and rational protein engineering.
Supplementary notes can be added here, including code, math, and images.